50 assays (manual) / 500 assays (microplate) / 450 assays (auto-analyser)
Prices exclude VAT
Available for shipping
| Content: | 50 assays (manual) / 500 assays (microplate) / 450 assays (auto-analyser) |
| Shipping Temperature: | Ambient |
| Storage Temperature: | Short term stability: 2-8oC, Long term stability: See individual component labels |
| Stability: | > 1 year under recommended storage conditions |
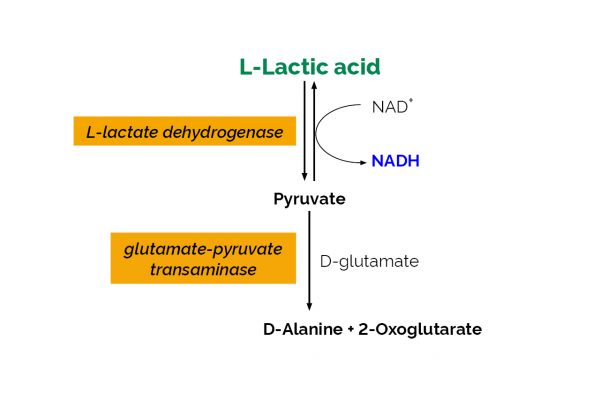
| Analyte: | L-Lactic Acid |
| Assay Format: | Spectrophotometer, Microplate, Auto-analyser |
| Detection Method: | Absorbance |
| Wavelength (nm): | 340 |
| Signal Response: | Increase |
| Linear Range: | 0.3 to 30 µg of L-lactic acid per assay |
| Limit of Detection: | 0.21 mg/L |
| Reaction Time (min): | ~ 10 min |
| Application examples: | Wine, beer, soft drinks, milk, dairy products (e.g. cream, milk / whey powder, cheese, condensed milk and yogurt), foods containing milk (e.g. dietetic foods, bakery products, baby food, chocolate, sweets and ice-cream), egg, egg products (e.g. egg powder), baking additives, vinegar, fruit and vegetables, processed fruit and vegetables (e.g. tomatoes), meat products, food additives, feed, paper (and cardboard), cosmetics, pharmaceuticals and other materials (e.g. biological cultures, samples, etc.). |
| Method recognition: | Methods based on this principle have been accepted by DIN, GOST, IDF, EEC, EN, ISO, OIV, IFU, AIJN and MEBAK |
The L-Lactic Acid (L-Lactate) Assay Kit is used for the specific measurement and analysis of L-lactic acid (L-lactate) in beverages, meat, dairy and food products.
Note for Content: The number of manual tests per kit can be doubled if all volumes are halved. This can be readily accommodated using the MegaQuantTM Wave Spectrophotometer (D-MQWAVE).
Explore our complete list of organic acid assay kits.

- Extended cofactors stability. Dissolved cofactors stable for > 1 year at 4oC.
- Very competitive price (cost per test)
- All reagents stable for > 2 years after preparation
- Rapid reaction
- Mega-Calc™ software tool is available from our website for hassle-free raw data processing
- Standard included
- Suitable for manual, microplate and auto-analyser formats
Megazyme “advanced” wine test kits general characteristics and validation.
Charnock, S. J., McCleary, B. V., Daverede, C. & Gallant, P. (2006). Reveue des Oenologues, 120, 1-5.
Many of the enzymatic test kits are official methods of prestigious organisations such as the Association of Official Analytical Chemicals (AOAC) and the American Association of Cereal Chemists (AACC) in response to the interest from oenologists. Megazyme decided to use its long history of enzymatic bio-analysis to make a significant contribution to the wine industry, by the development of a range of advanced enzymatic test kits. This task has now been successfully completed through the strategic and comprehensive process of identifying limitations of existing enzymatic bio-analysis test kits where they occurred, and then using advanced techniques, such as molecular biology (photo 1), to rapidly overcome them. Novel test kits have also been developed for analytes of emerging interest to the oenologist, such as yeast available nitrogen (YAN; see pages 2-3 of issue 117 article), or where previously enzymes were simply either not available, or were too expensive to employ, such as for D-mannitol analysis.
Hide AbstractGrape and wine analysis: Oenologists to exploit advanced test kits.
Charnock, S. C. & McCleary, B. V. (2005). Revue des Enology, 117, 1-5.
It is without doubt that testing plays a pivotal role throughout the whole of the vinification process. To produce the best possible quality wine and to minimise process problems such as “stuck” fermentation or troublesome infections, it is now recognised that if possible testing should begin prior to harvesting of the grapes and continue through to bottling. Traditional methods of wine analysis are often expensive, time consuming, require either elaborate equipment or specialist expertise and frequently lack accuracy. However, enzymatic bio-analysis enables the accurate measurement of the vast majority of analytes of interest to the wine maker, using just one piece of apparatus, the spectrophotometer (see previous issue No. 116 for a detailed technical review). Grape juice and wine are amenable to enzymatic testing as being liquids they are homogenous, easy to manipulate, and can generally be analysed without any sample preparation.
Hide AbstractRestricted O2 consumption in pea roots induced by hexanoic acid is linked to depletion of Krebs cycle substrates.
Casolo, V., Zancani, M., Pellegrini, E., Filippi, A., Gargiulo, S., Konnerup, D., Morandini, P. & Pedersen, O. (2023). Physiologia Plantarum, 175(5), e14024.
Plant roots are exposed to hypoxia in waterlogged soils, and they are further challenged by specific phytotoxins produced by microorganisms in such conditions. One such toxin is hexanoic acid (HxA), which, at toxic levels, causes a strong decline in root O2 consumption. However, the mechanism underlying this process is still unknown. We treated pea (Pisum sativum L.) roots with 20 mM HxA at pH 5.0 and 6.0 for a short time (1 h) and measured leakage of key electrolytes such as metal cations, malate, citrate and nonstructural carbohydrates (NSC). After treatment, mitochondria were isolated to assess their functionality evaluated as electrical potential and O2 consumption rate. HxA treatment resulted in root tissue extrusion of K+, malate, citrate and NSC, but only the leakage of the organic acids and NSC increased at pH 5.0, concomitantly with the inhibition of O2 consumption. The activity of mitochondria isolated from treated roots was almost unaffected, showing just a slight decrease in oxygen consumption after treatment at pH 5.0. Similar results were obtained by treating the pea roots with another organic acid with a short carbon chain, that is, butyric acid. Based on these results, we propose a model in which HxA, in its undissociated form prevalent at acidic pH, stimulates the efflux of citrate, malate and NSC, which would, in turn, cause starvation of mitochondrial respiratory substrates of the Krebs cycle and a consequent decline in O2 consumption. Cation extrusion would be a compensatory mechanism in order to restore plasma membrane potential.
Hide AbstractTreatment of food processing wastes for the production of medium chain fatty acids via chain elongation.
Battista, F., Zeni, A., Andreolli, M., Salvetti, E., Rizzioli, F., Lampis, S. & Bolzonella, D. (2024). Environmental Technology & Innovation, 33, 103453.
The production of medium chain fatty acids (MCFAs) through reverse β-oxidation was investigated both on synthetic and real substrates. From preliminary batch tests emerged that caproic acid was maximized under an acetate/ethanol molar ratio of 5:1 at neutral pH. This ratio was then adopted in different semi-continuous tests operating with different amounts of the two reactants. It emerged that the MCFAs yield reached the maximum level of 6.7% when the total molar substrate amount was around 40–45 mmol/d, while the process was inhibited for values higher than 400 mmol/d. Semi-continuous tests using real waste as substrates, namely food waste condensate, cheese whey, and winery wastewater, confirmed the results obtained with the synthetic substrates. Better performances were obtained when an adequate molar ratio of the acetate and the electron-donor compound was naturally present. Therefore, a MCFAs yield of 25% and 10.5% was obtained for condensate of food waste and acidic cheese whey, respectively. Regarding MCFAs composition, caproic acid was the dominant form but small concentrations of octanoic acid were also found in the tests where ethanol was the electron donor (synthetic substrates and food waste condensate). Octanoic acid was not produced in test where lactic acid represented the electron donor molecules (cheese whey). Condensate and synthetic samples were dominated by Pseudoclavibacter caeni with an abundance of 38.19% and 33.38% respectively, while Thomasclavelia (24.13%) and Caproiciproducens (11.68%) was the most representative genus in acidic cheese whey sample.
Hide AbstractMis-regulation of Zn and Mn homeostasis is a key phenotype of Cu stress in Streptococcus pyogenes.
Hong, Y., Mackenzie, E. S., Firth, S. J., Bolton, J. R., Stewart, L. J., Waldron, K. J. & Djoko, K. Y. (2023). Metallomics, 15(11), mfad064.
All bacteria possess homeostastic mechanisms that control the availability of micronutrient metals within the cell. Cross-talks between different metal homeostasis pathways within the same bacterial organism have been reported widely. In addition, there have been previous suggestions that some metal uptake transporters can promote adventitious uptake of the wrong metal. This work describes the cross-talk between Cu and the Zn and Mn homeostasis pathways in Group A Streptococcus (GAS). Using a ∆copA mutant strain that lacks the primary Cu efflux pump and thus traps excess Cu in the cytoplasm, we show that growth in the presence of supplemental Cu promotes downregulation of genes that contribute to Zn or Mn uptake. This effect is not associated with changes in cellular Zn or Mn levels. Co-supplementation of the culture medium with Zn or, to a lesser extent, Mn alleviates key Cu stress phenotypes, namely bacterial growth and secretion of the fermentation end-product lactate. However, neither co-supplemental Zn nor Mn influences cellular Cu levels or Cu availability in Cu-stressed cells. In addition, we provide evidence that the Zn or Mn uptake transporters in GAS do not promote Cu uptake. Together, the results from this study strengthen and extend our previous proposal that mis-regulation of Zn and Mn homeostasis is a key phenotype of Cu stress in GAS.
Hide AbstractMetformin Induces Lipogenesis and Apoptosis in H4IIE Hepatocellular Carcinoma Cells.
Park, D., Lee, S. & Boo, H. (2023). Development & Reproduction, 27(2), 77.
Metformin is the most widely used anti-diabetic drug that helps maintain normal blood glucose levels primarily by suppressing hepatic gluconeogenesis in type II diabetic patients. We previously found that metformin induces apoptotic death in H4IIE rat hepatocellular carcinoma cells. Despite its anti-diabetic roles, the effect of metformin on hepatic de novo lipogenesis (DNL) remains unclear. We investigated the effect of metformin on hepatic DNL and apoptotic cell death in H4IIE cells. Metformin treatment stimulated glucose consumption, lactate production, intracellular fat accumulation, and the expressions of lipogenic proteins. It also stimulated apoptosis but reduced autophagic responses. These metformin-induced changes were clearly reversed by compound C, an inhibitor of AMP-activated protein kinase (AMPK). Interestingly, metformin massively increased the production of reactive oxygen species (ROS), which was completely blocked by compound C. Metformin also stimulated the phosphorylation of p38 mitogen-activated protein kinase (p38MAPK). Finally, inhibition of p38MAPK mimicked the effects of compound C, and suppressed the metformin-induced fat accumulation and apoptosis. Taken together, metformin stimulates dysregulated glucose metabolism, intracellular fat accumulation, and apoptosis. Our findings suggest that metformin induces excessive glucose-induced DNL, oxidative stress by ROS generation, activation of AMPK and p38MAPK, suppression of autophagy, and ultimately apoptosis.
Hide AbstractDevelopment of In Situ Product Recovery (ISPR) System Using Amberlite IRA67 for Enhanced Biosynthesis of Hyaluronic Acid by Streptococcus zooepidemicus.
Abdullah Thaidi, N. I., Mohamad, R., Wasoh, H., Kapri, M. R., Ghazali, A. B., Tan, J. S., Rios-Solis, L. & Halim, M. (2023). Life, 13(2), 558.
High broth viscosity due to the accumulation of hyaluronic acid (HA) causes a limited yield of HA. It is a major problem of HA production using Streptococcus zooepidemicus. Extractive fermentation via in situ product recovery (ISPR) was utilized to enhance the HA production. Resins from Amberlite: IRA400 Cl; IRA900 Cl; IRA410 Cl; IRA402 Cl; and IRA67 were tested for the HA adsorption. IRA67 showed high adsorption capacity on HA. The study of the adsorption via a 2 L stirred tank bioreactor of S. zooepidemicus fermentation was investigated to elucidate the adsorption of HA onto IRA67 in dispersed and integrated internal column systems. The application of a dispersed IRA67 improved the HA production compared to the fermentation without resin addition by 1.37-fold. The HA production was further improved by 1.36-fold with an internal column (3.928 g/L) over that obtained with dispersed IRA67. The cultivation with an internal column shows the highest reduction of viscosity value after the addition of IRA67 resin: from 58.8 to 23.7 (mPa·s), suggesting the most effective ISPR of HA. The improved biosynthesis of HA indicated that an extractive fermentation by ISPR adsorption is effective and may streamline the HA purification.
Hide AbstractAntioxidant and Functional Features of Pre-Fermented Ingredients Obtained by the Fermentation of Milling By-Products.
Siroli, L., Giordani, B., Rossi, S., Gottardi, D., McMahon, H., Augustyniak, A., Menon, A., Vannini, L., Vitali, B., Patrignan, F. & Lanciotti, R. (2022). Fermentation, 8(12), 722.
The use of milling by-products as ingredients in food formulations has increased gradually over the past years, due to their well-recognized health properties. Fermentation performed with selected microbial strains or microbial consortia is the most promising way to reduce antinutritional factors of cereals and bran, while increasing their nutritional and functional properties. This work, developed within the BBI project INGREEN, was aimed to study the functional, nutritional and technological features of a pre-fermented ingredient obtained from the fermentation of a mixture of rye bran and wheat germ by a selected microbial consortium composed of yeasts (Kazachstania unispora and Kazachstania servazii) and lactic acid bacteria (Latilactobacillus curvatus) using as reference the unfermented mixture and the same mixture fermented by a baker’s yeast. The selected microbial consortium improved the complexity of the volatile molecules such as acids, alcohols and esters. A better retention of color parameters was maintained compared to the product fermented by a baker’s yeast. In addition, the fermentation by the selected consortium showed a significant increase in short chain fatty acids (more than 5-fold), antioxidant activity (22–24%), total phenol content (53-71%), bioactive peptides (39-52%), a reduction of 20-28% in phytic acid content and an increase in prebiotic activity not only compared to the unfermented product but also compared to the preferment obtained with a baker’s yeast. Overall, the fermentation by the selected microbial consortium can be considered a valuable way to valorize milling by-products and promote their exploitation as food ingredients.
Hide AbstractMetabolic reprogramming of OPA1-deficient cells.
Dai, W., Wang, Z., Wang, Q. A., Chan, D. & Jiang, L. (2022). Cell Mol. Life Sci., 79(10), 517.
OPA1, a dynamin-related GTPase mutated in autosomal dominant optic atrophy, is essential for the fusion of the inner mitochondrial membrane. Although OPA1 deficiency leads to impaired mitochondrial morphology, the role of OPA1 in central carbon metabolism remains unclear. Here, we aim to explore the functional role and metabolic mechanism of OPA1 in cell fitness beyond the control of mitochondrial fusion. We applied [U-13C]glucose and [U-13C]glutamine isotope tracing techniques to OPA1-knockout (OPA1-KO) mouse embryonic fibroblasts (MEFs) compared to OPA1 wild-type (OPA1-WT) controls. Furthermore, the resulting tracing data were integrated by metabolic flux analysis to understand the underlying metabolic mechanism through which OPA1 deficiency reprograms cellular metabolism. OPA1-deficient MEFs were depleted of intracellular citrate, which was consistent with the decreased oxygen consumption rate in these cells with mitochondrial fission that is not balanced by mitochondrial fusion. Whereas oxidative glucose metabolism was impaired, OPA1-deficient cells activated glutamine-dependent reductive carboxylation and subsequently relied on this reductive metabolism to produce cytosolic citrate as a predominant acetyl-CoA source for de novo fatty acid synthesis. Prevention of cytosolic glutamine reductive carboxylation by GSK321, an inhibitor of isocitrate dehydrogenase 1 (IDH1), largely repressed lipid synthesis and blocked cell proliferation in OPA1-deficient MEFs. Our data support that, when glucose oxidation failed to support lipogenesis and proliferation in cells with unbalanced mitochondrial fission, OPA1 deficiency stimulated metabolic anaplerosis into glutamine-dependent reductive carboxylation in an IDH1-mediated manner.
Hide AbstractEffect of inoculation strategy with autochthonous Oenococcus oeni strains on aroma development in Rioja Alavesa Tempranillo wines.
Diez-Ozaeta, I., Lavilla, M. & Amárita, F. (2022). LWT, 162, 113399.
The potential use as malolactic starters of four indigenous strains of Oenococcus oeni was evaluated under different inoculation regimes. Among others, the fermentative capacity of strains, their degree of implantation, the main oenological parameters as well as their ability to modulate the aromatic profile of wines, were analysed. Main results elucidated that co-inoculation led to the prompt consecution of malolactic fermentation (MLF), highlighting the performance of indigenous Oenococcus oeni P2A strain and commercial O. oeni Viniflora OENOS strain which finished the process 20-30 days earlier compared to batches that undergo sequential inoculation. Moreover, inoculation strategy did also have an important influence on the volatile profile of wines. Co-inoculated wines significantly showed less concentration of volatile compounds. Main reduction was detected in higher alcohols and acids. Lower concentration of acids and higher alcohols may prevent the masking of desired aroma attributes. Indeed, in co-inoculated wines, the perception of ripe fruit aroma was highlighted over the others, and was extensively perceived by panellists in comparison with their respective sequentially inoculated wines. Above all, it was elucidated the suitability of the strain P2A, resulting an advantageous alternative to significantly reduce the overall winemaking time as well as to better control the fermentative process.
Hide AbstractResource recovery from the anaerobic digestion of food waste is underpinned by cross-kingdom microbial activities.
Nzeteu, C., Joyce, A., Thorn, C., McDonnell, K., Shirran, S., O'Flaherty, V. & Abram, F. (2021). Bioresource Technology Reports, 16, 100847.
As the human population grows on the planet so does the generation of waste and particularly that of food waste. In order to tackle the world sustainability crisis, efforts to recover products from waste are critical. Here, we anaerobically recovered volatile fatty acids (VFAs) from food waste and analysed the microbial populations underpinning the process. An increased contribution of fungi relative to bacteria was observed throughout the reactor operation, with both kingdoms implicated into the main three steps of anaerobic digestion occurring within our systems: hydrolysis, acidogenesis and acetogenesis. Overall, Ascomycota, Proteobacteria and Firmicutes were found to drive the anaerobic digestion of food waste, with butyrate as the most abundant VFA likely produced by Clostridium using lactate as a precursor. Taken together we demonstrate that the generation of products of added-value from food waste results from cross-kingdoms microbial activities implicating fungi and bacteria.
Hide AbstractHighly Selective and Label-Free AuNPs Based Optical Sensor Development Significant in Smartphone Sensing of L-Lactate in Food Samples.
Rattu, G. & Krishna, P. M. (2021). IEEE Transactions on Nanotechnology, 20, 635-643.
L-lactate is an essential organic chemical in food processing, clinical, chemical, and fermentation industries. AuNPs were synthesized by chemical reduction reaction method and functionalized with 3-aminophenylboronic acid (3-APBA). The structural, morphological, FTIR, DLS, zeta potential characterization studies were performed to optimize the sensing mechanism. First time reporting facile enzyme-free sensor for the detection of L-lactate content in milk. Initially, the sensor has been optimized for nano reagent concentration, pH and time. 3-APBA@AuNPs showed a stable SPR intensity and forms a reversible tetrahedral ester complex with the analyte L-lactate (100 mM) resulting the NPs aggregation and revealing the blue shift of UV-VIS absorption wavelength (λ = 300 nm to 288 nm). Nestle- Lactogen, an India commercial milk powder has been used for the standardization study of lactate sensing. The milk samples spiked with L-Lactate (1-20mM) showed an increase in UV-VIS absorption peak intensity with concentration, and estimated the lower detection limit (LOD) = 1.1 mM using the calibration graphs. The optical sensing mechanism is explained may be due to π-π* transition of AuNPs stabilized by tetragonal complex of L-lactate bonded to 3-boronate via nucleophilic attack on boron atom. The nanosensor data was validated with commercial Megazyme kit and conventional HPLC technique. In commercial product Amul-India milk, the L-lactate value estimated as 0.678Mm with 95% correlation. Hence, the proposed sensor technique could be an alternate to the high cost conventional methods and at next level, it can be interfaced to smart phone to develop an onsite food, pharma and clinical device applications.
Hide AbstractNMS-873 functions as a dual inhibitor of mitochondrial oxidative phosphorylation.
Bouwer, M. F., Hamilton, K. E., Jonker, P. B., Kuiper, S. R., Louters, L. L. & Looyenga, B. D. (2021). Biochimie, 185, 33-42.
Small-molecule inhibitors of enzyme function are critical tools for the study of cell biological processes and for treatment of human disease. Identifying inhibitors with suitable specificity and selectivity for single enzymes, however, remains a challenge. In this study we describe our serendipitous discovery that NMS-873, a compound that was previously identified as a highly selective allosteric inhibitor of the ATPase valosin-containing protein (VCP/p97), rapidly induces aerobic fermentation in cultured human and mouse cells. Our further investigation uncovered an unexpected off-target effect of NMS-873 on mitochondrial oxidative phosphorylation, specifically as a dual inhibitor of Complex I and ATP synthase. This work points to the need for caution regarding the interpretation of cell survival data associated with NMS-873 treatment and indicates that cellular toxicity associated with its use may be caused by both VCP/p97-dependent and VCP/p97-independent mechanisms.
Hide AbstractChemical Composition of Sour Beer Resulting from Supplementation the Fermentation Medium with Magnesium and Zinc Ions.
Ciosek, A., Fulara, K., Hrabia, O., Satora, P. & Poreda, A. (2020). Biomolecules, 10(12), 1599.
The bioavailability of minerals, such as zinc and magnesium, has a significant impact on the fermentation process. These metal ions are known to influence the growth and metabolic activity of yeast, but there are few reports on their effects on lactic acid bacteria (LAB) metabolism during sour brewing. This study aimed to evaluate the influence of magnesium and zinc ions on the metabolism of Lactobacillus brevis WLP672 during the fermentation of brewers’ wort. We carried out lactic acid fermentations using wort with different mineral compositions: without supplementation; supplemented with magnesium at 60 mg/L and 120 mg/L; and supplemented with zinc at 0.4 mg/L and 2 mg/L. The concentration of organic acids, pH of the wort and carbohydrate use was determined during fermentation, while aroma compounds, real extract and ethanol were measured after the mixed fermentation. The addition of magnesium ions resulted in the pH of the fermenting wort decreasing more quickly, an increase in the level of L-lactic acid (after 48 h of fermentation) and increased concentrations of some volatile compounds. While zinc supplementation had a negative impact on the L. brevis strain, resulting in a decrease in the L-lactic acid content and a higher pH in the beer. We conclude that zinc supplementation is not recommended in sour beer production using L. brevis WLP672.
Hide AbstractSingle-use printed biosensor for l-lactate and its application in bioprocess monitoring.
Theuer, L., Randek, J., Junne, S., Neubauer, P., Mandenius, C. F. & Beni, V. (2020). Processes, 8(3), 321.
There is a profound need in bioprocess manufacturing for low-cost single-use sensors that allow timely monitoring of critical product and production attributes. One such opportunity is screen-printed enzyme-based electrochemical sensors, which have the potential to enable low-cost online and/or off-line monitoring of specific parameters in bioprocesses. In this study, such a single-use electrochemical biosensor for lactate monitoring is designed and evaluated. Several aspects of its fabrication and use are addressed, including enzyme immobilization, stability, shelf-life and reproducibility. Applicability of the biosensor to off-line monitoring of bioprocesses was shown by testing in two common industrial bioprocesses in which lactate is a critical quality attribute (Corynebacterium fermentation and mammalian Chinese hamster ovary (CHO) cell cultivation). The specific response to lactate of the screen-printed biosensor was characterized by amperometric measurements. The usability of the sensor at typical industrial culture conditions was favorably evaluated and benchmarked with commonly used standard methods (HPLC and enzymatic kits). The single-use biosensor allowed fast and accurate detection of lactate in prediluted culture media used in industrial practice. The design and fabrication of the biosensor could most likely be adapted to several other critical bioprocess analytes using other specific enzymes. This makes this single-use screen-printed biosensor concept a potentially interesting and versatile tool for further applications in bioprocess monitoring.
Hide Abstract