100 assays (50 of each) per kit
Prices exclude VAT
Available for shipping
| Content: | 100 assays (50 of each) per kit |
| Shipping Temperature: | Ambient |
| Storage Temperature: | Short term stability: 2-8oC, Long term stability: See individual component labels |
| Stability: | > 1 year under recommended storage conditions |
| Analyte: | Ammonia, Nitrogen, Urea, YAN |
| Assay Format: | Spectrophotometer, Auto-analyser |
| Detection Method: | Absorbance |
| Wavelength (nm): | 340 |
| Signal Response: | Decrease |
| Linear Range: | 0.2 to 7 µg of ammonia (0.3 to 14 µg of urea) per assay |
| Limit of Detection: | 0.13 mg/L (urea), 0.07 mg/L (ammonia) |
| Reaction Time (min): | ~ 10 min |
| Application examples: | Wine, grape juice, must, fruit juices, soft drinks, milk, cheese, meat, processed meat, bakery products, seafood, fertilizers, feed, pharmaceuticals, cosmetics, water (e.g. swimming-pool water), Kjeldahl analysis, paper (and cardboard) and other materials (e.g. biological cultures, samples, etc.). |
| Method recognition: | Methods based on this principle have been accepted by NEN and MEBAK |
The Urea/Ammonia (Rapid) test kit is suitable for the specific and rapid measurement and analysis of urea and ammonia in water, beverages, milk and food products.
Note for Content: The number of manual tests per kit can be doubled if all volumes are halved. This can be readily accommodated using the MegaQuantTM Wave Spectrophotometer (D-MQWAVE).
Other nitrogen assay kits also available.

- Extended cofactors stability. Dissolved cofactors stable for > 1 year at 4oC.
- Very rapid reaction due to use of uninhibited glutamate dehydrogenase
- Enzymes supplied as stable Suspensions
- Very competitive price (cost per test)
- All reagents stable for > 2 years after preparation
- Mega-Calc™ software tool is available from our website for hassle-free raw data processing
- Standard included
Megazyme “advanced” wine test kits general characteristics and validation.
Charnock, S. J., McCleary, B. V., Daverede, C. & Gallant, P. (2006). Reveue des Oenologues, 120, 1-5.
Many of the enzymatic test kits are official methods of prestigious organisations such as the Association of Official Analytical Chemicals (AOAC) and the American Association of Cereal Chemists (AACC) in response to the interest from oenologists. Megazyme decided to use its long history of enzymatic bio-analysis to make a significant contribution to the wine industry, by the development of a range of advanced enzymatic test kits. This task has now been successfully completed through the strategic and comprehensive process of identifying limitations of existing enzymatic bio-analysis test kits where they occurred, and then using advanced techniques, such as molecular biology (photo 1), to rapidly overcome them. Novel test kits have also been developed for analytes of emerging interest to the oenologist, such as yeast available nitrogen (YAN; see pages 2-3 of issue 117 article), or where previously enzymes were simply either not available, or were too expensive to employ, such as for D-mannitol analysis.
Hide AbstractGrape and wine analysis: Oenologists to exploit advanced test kits.
Charnock, S. C. & McCleary, B. V. (2005). Revue des Enology, 117, 1-5.
It is without doubt that testing plays a pivotal role throughout the whole of the vinification process. To produce the best possible quality wine and to minimise process problems such as “stuck” fermentation or troublesome infections, it is now recognised that if possible testing should begin prior to harvesting of the grapes and continue through to bottling. Traditional methods of wine analysis are often expensive, time consuming, require either elaborate equipment or specialist expertise and frequently lack accuracy. However, enzymatic bio-analysis enables the accurate measurement of the vast majority of analytes of interest to the wine maker, using just one piece of apparatus, the spectrophotometer (see previous issue No. 116 for a detailed technical review). Grape juice and wine are amenable to enzymatic testing as being liquids they are homogenous, easy to manipulate, and can generally be analysed without any sample preparation.
Hide AbstractExperimental study and modelling of a packed bed bioreactor for urea removal in wines.
Mazzù, R., Tavilli, E. & Fidaleo, M. (2023). Food and Bioproducts Processing, 140, 230-241.
The study involved the development and modeling of a fixed-bed bioreactor for the removal of urea from wines. The reactor, based on the immobilization of acid urease enzyme, was studied under both stationary and non-stationary conditions. The developed model, including internal and external catalyst particle mass transfer, Michaelis-Menten kinetics, convection and dispersion in the liquid along the reactor axis, was able to produce urea concentration profiles in both the solid and liquid phases under various volumetric flow rates and inlet urea concentrations. The experimental results were in good agreement with the model predictions, the mean relative error between simulated and experimental outlet ammonia concentration ranging from 4.1 % to 16.4 %. Model simulations confirmed that in wines the reaction kinetics is of the pseudo-first order and that internal and external catalyst particle diffusion limitations are negligeable. Simulations of the decrease of urea concentration as a function of space velocity for the reactor under study operating in the continuous mode and for three different wines were obtained confirming that urea removal by immobilized urease in wines is more difficult than in sake. The results obtained form the basis for the designing and scaling up of bioreactors for the treatment of wines.
Hide AbstractScheffersomyces stipitis ability to valorize different residual biomasses for vitamin B9 production.
Mastella, L., Senatore, V., Beltrani, T. & Branduardi, P. (2022). Microbial Biotechnology.
Sugar beet pulp (SBP), sugar beet molasses (SBM) and unfermented grape marcs (UGM) represent important waste in the agro-food sector. If suitably pre-treated, hexose and pentose sugars can be released in high quantities and can subsequently be used by appropriate cell factories as growth media and for the production of (complex) biomolecules, accomplishing the growing demand for products obtained from sustainable resources. One example is vitamin B9 or folate, a B-complex vitamin currently produced by chemical synthesis, almost exclusively in the oxidized form of folic acid (FA). It is therefore desirable to develop novel competitive strategies for replacing its current fossil-based production with a sustainable bio-based process. In this study, we assessed the production of natural folate by the yeast Scheffersomyces stipitis, investigating SBM, SBP and UGM as potential growth media. Pre-treatment of SBM and SBP had previously been optimized in our laboratory; thus, here we focused only on UGM pre-treatment and hydrolysis strategies for the release of fermentable sugars. Then, we optimized the growth of S. stipitis on the three media formulated from those biomasses, working on inoculum pre-adaptation, oxygen availability and supplementation of necessary nutrients to support the microorganism. Folate production, measured with a microbiological assay, reached 188.2 ± 24.86 μg/L on SBM, 130.6 ± 1.34 μg/L on SBP and 101.9 ± 6.62 μg/L on UGM. Here, we demonstrate the flexibility of S. stipitis in utilizing different residual biomasses as growth media. Moreover, we assessed the production of folate from waste, and to the best of our knowledge, we obtained the highest production of folate from residual biomasses ever reported, providing the first indications for the future development of this microbial production process.
Hide AbstractStreamlining of a synthetic co‐culture towards an individually controllable one‐pot process for polyhydroxyalkanoate production from light and CO2.
Kratzl, F., Kremling, A. & Pflüger‐Grau, K. Engineering in Life Sciences, (2022), In Press.
Rationally designed synthetic microbial consortia carry a vast potential for biotechnological applications. The application of such a consortium in a bioprocess, however, requires tight and individual controllability of the involved microbes. Here, we present the streamlining of a co-cultivation process consisting of Synechococcus elongatus cscB and Pseudomonas putida for the production of polyhydroxyalkanoates (PHA) from light and CO2. First, the process was improved by employing P. putida cscRABY, a strain with a higher metabolic activity towards sucrose. Next, the individual controllability of the co-culture partners was addressed by providing different nitrogen sources, each exclusively available for one strain. By this, the growth rate of the co-culture partners could be regulated individually, and defined conditions could be set. The molC/molN ratio, a key value for PHA accumulation, was estimated from the experimental data, and the necessary feeding rates to obtain a specific ratio could be predicted. This information was then implemented in the co-cultivation process, following the concept of a DBTL-cycle. In total, the streamlining of the process resulted in an increased maximal PHA titer of 393 mg/L and a PHA production rate of 42.1 mg/(L.day).
Hide AbstractPerformance evaluation of an enzymatic spectrophotometric method for milk urea nitrogen.
Portnoy, M., Coon, C. & Barbano, D. M. (2021). Journal of Dairy Science.
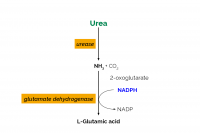
Our objective was to determine the within and between laboratory performance of an enzymatic spectrophotometric method for milk urea nitrogen (MUN) determination. This method first uses urease to hydrolyze urea into ammonia and carbon dioxide. Next, ammonia (as ammonium ions) reacts with 2-oxoglutarate, in the presence of reduced nicotinamide-adenine dinucleotide phosphate (NADPH) and the enzyme glutamate dehydrogenase (GlDH), to form l-glutamic acid, water, and NADP+. The change in light absorption at 340 nm due to the conversion of NADPH to NADP+ is stoichiometrically a function of the MUN content of a milk sample. The relative within (RSDr) and between (RSDR) laboratory method performance values for the MUN enzymatic spectrophotometric method were 0.57% and 0.85%, respectively, when testing individual farm milks. The spectrophotometric MUN method demonstrated better within and between laboratory performance than the International Dairy Federation differential pH MUN method with a much lower RSDr (0.57 vs. 1.40%) and RSDR (0.85 vs. 4.64%). The spectrophotometric MUN method also had similar method performance statistics as other AOAC International official validated chemical methods for primary milk component determinations, with the average of all RSDr and RSDR values being <1%. An official collaborative study of the enzymatic spectrophotometric MUN method is needed to achieve International Dairy Federation, AOAC International, and International Organization for Standardization official method status.
Hide AbstractEvaluation of Dietary Administration of Chestnut and Quebracho Tannins on Growth, Serum Metabolites and Fecal Parameters of Weaned Piglets.
Caprarulo, V., Hejna, M., Giromini, C., Liu, Y., Dell’Anno, M., Sotira, S., Reggi, S., Sgoifo-Rossi, C. A., Callegari, M. L. & Rossi, L. (2020). Animals, 10(11), 1945.
In pig livestock, alternatives to in-feed antibiotics are needed to control enteric infections. Plant extracts such as tannins can represent an alternative as a natural source of functional compounds. The aim of this study was to evaluate the in vitro digestibility and in vivo effects of oral supplementation of combined chestnut (Ch) and quebracho (Qu) tannins in order to establish if they can induce a positive effect on weaned piglets’ performance, metabolic status and fecal parameters. In vitro digestibility (dry matter, DM) of diets was calculated using a multi-step enzymatic technique. In vitro digested diet samples were further tested on an intestinal porcine enterocyte cell line (IPEC-J2). Weaned piglets (n = 120; 28 ± 2 day old) were randomly allotted to two groups (12 pens in total with 10 pigs per pen): control (Ctrl) and treatment (Ch/Qu). After one week of adaptation (day 0), 35-day-old piglets in the Ctrl group were fed a Ctrl diet and the Ch/Qu group were fed with 1.25% Ch/Qu for 40 days. Body weight and feed intake per pen were recorded weekly. At day 40, blood and fecal samples were collected. Principal metabolic parameters were evaluated from blood samples by enzymatic colorimetric analysis. Total phenolic compounds, urea, and ammonia in feces were analyzed (Megazyme International, Bray, Ireland). In vitro digestibility and cell viability assays showed that the inclusion of 1.25% Ch/Qu slightly reduced diet digestibility compared with the Ctrl diet, while intestinal cell viability was not altered with low concentrations of Ch/Qu digesta compared with Ctrl. In vivo results did not show any adverse effects of Ch/Qu on feed intake and growth performance, confirming that dietary inclusion of Ch/Qu at a concentration of 1.25% did not impair animal performance. The decreased diet DM digestibility in the Ch/Qu diet may cause increased serum concentration of albumin (Ctrl: 19.30 ± 0.88; Ch/Qu: 23.05 ± 0.88) and albumin/globulin ratio (Ctrl: 0.58 ± 0.04; Ch/Qu: 0.82 ± 0.04), but decreased creatinine (Ctrl: 78.92 ± 4.18; Ch/Qu: 54.82 ± 4.18) and urea (Ctrl: 2.18 ± 0.19; Ch/Qu: 0.95 ± 0.19) compared with Ctrl. Pigs in the Ch/Qu group contained higher (p < 0.05) concentrations of fecal phenolic compounds and nitrogen than the Ctrl group, while fecal ammonia and urea were not affected by tannins. In conclusion, Ch/Qu tannin supplementation did not influence growth performance. Although lower digestibility was observed in the diet supplemented with Ch/Qu tannins, Ch/Qu supplementation did not show any adverse effect on intestinal epithelial cell viability.
Hide AbstractFlow-based method for the determination of biomarkers urea and ammoniacal nitrogen in saliva.
Thepchuay, Y., Costa, C. F., Mesquita, R. B., Sampaio-Maia, B., Nacapricha, D. & Rangel, A. O. (2020). Bioanalysis, 12(7), 455-465.
Aim: Salivary urea and ammonium levels are potential biomarkers for chronic kidney disease. A fast and efficient assessment of these compounds in the saliva of healthy and diseased individuals may be a useful tool to monitor kidney function. Materials & methods: Ammonium ions were measured with an ammonia selective electrode after conversion to ammonia gas. A urease reactor was incorporated in the manifold to hydrolyze urea to ammonium, thereby providing values of ammonia from both urea and ammonium ions in the sample. The accuracy of the method was assessed by comparison with a commercially available kit for urea and ammonium determination. Conclusion: A sequential injection method for the biparametric determination of salivary urea and ammonium employing a single sequential injection manifold was successfully applied to samples collected from both healthy volunteers and chronic kidney disease patients.
Hide AbstractDifferential cytokine and metabolite production by cervicovaginal epithelial cells infected with Lactobacillus crispatus and Ureaplasma urealyticum.
Cavanagh, M., Amabebe, E. & Anumba, D. O. (2020). Anaerobe, 62, 102101.
Introduction: We sought to quantify targeted metabolites (d-lactate, pyruvate, urea, ammonia) and the cytokine IL-8 produced by human cervicovaginal epithelial cells co-cultured with Ureaplasma urealyticum (a preterm birth-associated bacterium) or Lactobacillus crispatus (a healthy vaginal commensal associated with term birth). Methods: Concentrations of D-lactate, pyruvate, urea and ammonia measured by enzyme-based spectrophotometry and IL-8 by ELISA were determined and compared between monolayer-cultured HeLa cells (ATCC 35241) infected with strains of U. urealyticum (ATCC 27618, 0.5 mL = 3640 CFU/mL, U. urealyticum) or L. crispatus (ATCC 33820, MOI = 10,000, 1000 and 100, L. crispatus) and incubated in 5% CO2 at 37°C for 24 h. Uninfected HeLa cells (Hc) were used as controls and cytotoxicity was determined by the amount (optical density) of lactate dehydrogenase (LDH) released by the dead HeLa cells. Results: The amount of LDH released by untreated Hc (P = 0.002) and U. urealyticum-infected cells (P < 0.0001) was higher than those of L. crispatus-infected cells, with U. urealyticum-infected cells recording the highest % cytotoxicity and L. crispatus-infected cells MOI 10,000 (Lc10,000) the least (P < 0.0001). Though there was no significant difference in the concentration of urea between the samples, U. urealyticum-infected cells showed higher ammonia compared to other samples (p = 0.03). In contrast, all L. crispatus samples had higher D-lactate than untreated Hc (p = 0.01) and U. urealyticum-infected cells (P = 0.01). Also, Lc10,000 had the highest D-lactate (p = 0.001) and lowest pyruvate (P = 0.04, excluding UU) compared to other samples. Furthermore, U. urealyticum-infected cells produced the highest IL-8 (P = 0.01) compared to other samples, with Lc10,000 producing undetectable levels. Conclusion: Infection of cervicovaginal epithelial cells by U. urealyticum stimulates production of ammonia from urea and induces elevated IL-8 production possibly leading to significantly higher cytotoxicity. In contrast, L. crispatus appeared protective against HeLa cell inflammation and death, producing more D-lactate and less IL-8, consistent with a role for L. crispatus in promoting vaginal floral health and reducing infection/inflammation-associated preterm birth.
Hide AbstractCheah, W. Y., Show, P. L., Juan, J. C., Chang, J. S. & Ling, T. C. (2018). Energy Conversion and Management, 164, 188-197.
Microalgae are a promising feedstock for biofuel generation. Economical and effective mass cultivation is essential for greater feasibility in microalgal-based biofuel full applications. The present study reported on cultivation of Chlorella sorokiniana CY-1 in palm oil mill effluent (POME) under photoautotrophic and mixotrophic cultivation. Enhancement of biomass and lipid productions were carried out by using glucose, urea and glycerol supplementations. Mixotrophic cultivation was more effective than photoautotrophic condition. Glycerol addition exhibited greater microalgae growth performance compared to supplementing glucose or urea. Biomass (1.68 g L-1) and lipid (15.07%) production were highest in POME medium with combinations of 200 mg L-1 urea, glucose and glycerol supplementation. Chlorella sorokiniana CY-1 grown in POME with glucose and glycerol supplementation gave considerably comparable yields as in all supplements-added POME medium. Ideal fatty acids compositions shown in urea and glycerol supplemented-POME medium though lower biomass production obtained. The pollutant remediation efficiencies attained were 63.85% COD, 91.54% TN and 83.25% TP in all supplements-added medium. The estimated net energy ratio was 0.55 and nutrient cost could be reduced up to 76%. Cheap and effective carbon and nutrients supplementation is essential to minimize the economic impact and maximize yields in commercial scale microalgae cultivation for biofuel production and environmental sustainability.
Hide AbstractLi, Y., Zhang, Y., Hussey, N. E. & Dai, X. (2016). Rapid Communications in Mass Spectrometry, 30(1), 1-8.
Rationale: Stable isotope analysis (SIA) provides a powerful tool to investigate diverse ecological questions for marine species, but standardized values are required for comparative assessments. For elasmobranchs, their unique osmoregulatory strategy involves retention of 15N -depleted urea in body tissues and this may bias δ15N values. This may be a particular problem for large predatory species, where δ15N discrimination between predator and consumed prey can be small. Methods: We evaluated three treatments (deionized water rinsing [DW], chloroform/methanol [LE] and combined chloroform/methanol and deionized water rinsing [LE+DW]) applied to white muscle tissue of 125 individuals from seven pelagic shark species to (i) assess urea and lipid effects on stable isotope values determined by IRMS and (ii) investigate mathematical normalization of these values. Results: For all species examined, the δ15N values and C:N ratios increased significantly following all three treatments, identifying that urea removal is required prior to SIA of pelagic sharks. The more marked change in δ15N values following DW (1.3 ± 0.4‰) and LE+DW (1.2 ± 0.6‰) than following LE alone (0.7 ± 0.4‰) indicated that water rinsing was more effective at removing urea. The DW and LE+DW treatments lowered the %N values, resulting in an increase in C:N ratios from the unexpected low values of 13N values of all species also increased significantly following LE and LE+DW treatments. Conclusions: Given the mean change in δ15N (1.2 ± 0.6‰) and δ13N values (0.7 ± 0.4‰) across pelagic shark species, it is recommended that muscle tissue samples be treated with LE+DW to efficiently extract both urea and lipids to standardize isotopic values. Mathematical normalization of urea and lipid-extracted δ15NLE+DW and δ13CLE+DW values using the lipid-extracted δ15NLE and δ13CLE data were established for all pelagic shark species.
Hide AbstractSlininger, P. J., Dien, B. S., Kurtzman, C. P., Moser, B. R., Bakota, E. L., Thompson, S. R., O'Bryan, P. J., Cotta, M. A., Balan, V., Jin, M., Sousa, L. D. C. & Dale, B. E. & Sousa, L. D. C. (2016). Biotechnology and Bioengineering, 113(8), 1676-1690.
Oleaginous yeasts can convert sugars to lipids with fatty acid profiles similar to those of vegetable oils, making them attractive for production of biodiesel. Lignocellulosic biomass is an attractive source of sugars for yeast lipid production because it is abundant, potentially low cost, and renewable. However, lignocellulosic hydrolyzates are laden with byproducts which inhibit microbial growth and metabolism. With the goal of identifying oleaginous yeast strains able to convert plant biomass to lipids, we screened 32 strains from the ARS Culture Collection, Peoria, IL to identify four robust strains able to produce high lipid concentrations from both acid and base-pretreated biomass. The screening was arranged in two tiers using undetoxified enzyme hydrolyzates of ammonia fiber expansion (AFEX)-pretreated cornstover as the primary screening medium and acid-pretreated switch grass as the secondary screening medium applied to strains passing the primary screen. Hydrolyzates were prepared at ~18-20% solids loading to provide ~110 g/L sugars at ~56:39:5 mass ratio glucose:xylose:arabinose. A two stage process boosting the molar C:N ratio from 60 to well above 400 in undetoxified switchgrass hydrolyzate was optimized with respect to nitrogen source, C:N, and carbon loading. Using this process three strains were able to consume acetic acid and nearly all available sugars to accumulate 50–65% of cell biomass as lipid (w/w), to produce 25-30 g/L lipid at 0.12-0.22 g/L/h and 0.13-0.15 g/g or 39-45% of the theoretical yield at pH 6 and 7, a performance unprecedented in lignocellulosic hydrolyzates. Three of the top strains have not previously been reported for the bioconversion of lignocellulose to lipids. The successful identification and development of top-performing lipid-producing yeast in lignocellulose hydrolyzates is expected to advance the economic feasibility of high quality biodiesel and jet fuels from renewable biomass, expanding the market potential for lignocellulose-derived fuels beyond ethanol for automobiles to the entire U.S. transportation market.
Hide AbstractMilne, N., Luttik, M. A. H., Rojas, H. C., Wahl, A., Van Maris, A. J. A., Pronk, J. T. & Daran, J. M. (2015). Metabolic Engineering, 30, 130-140.
In microbial processes for production of proteins, biomass and nitrogen-containing commodity chemicals, ATP requirements for nitrogen assimilation affect product yields on the energy producing substrate. In Saccharomyces cerevisiae, a current host for heterologous protein production and potential platform for production of nitrogen-containing chemicals, uptake and assimilation of ammonium requires 1 ATP per incorporated NH3. Urea assimilation by this yeast is more energy efficient but still requires 0.5 ATP per NH3 produced. To decrease ATP costs for nitrogen assimilation, the S. cerevisiae gene encoding ATP-dependent urease (DUR1,2) was replaced by a Schizosaccharomyces pombe vgene encoding ATP-independent urease (ure2), along with its accessory genes ureD, ureF and ureG. Since S. pombe ure2 is a Ni2+-dependent enzyme and Saccharomyces cerevisiae does not express native Ni2+-dependent enzymes, the S. pombe high-affinity nickel-transporter gene (nic1) was also expressed. Expression of the S. pombe genes into dur1,2 Δ S. cerevisiae yielded an in vitro ATP-independent urease activity of 0.44±0.01 µmol min-1 mg protein-1 and restored growth on urea as sole nitrogen source. Functional expression of the Nic1 transporter was essential for growth on urea at low Ni2+ concentrations. The maximum specific growth rates of the engineered strain on urea and ammonium were lower than those of a DUR1,2 reference strain. In glucose-limited chemostat cultures with urea as nitrogen source, the engineered strain exhibited an increased release of ammonia and reduced nitrogen content of the biomass. Our results indicate a new strategy for improving yeast-based production of nitrogen-containing chemicals and demonstrate that Ni2+-dependent enzymes can be functionally expressed in S. cerevisiae.
Hide AbstractNaert, L., Vande Vyvere, B., Verhoeven, G., Duchateau, L., De Smet, S. & Coopman, F. (2013). Vlaams Diergeneeskundig Tijdschrift, 82(1), 23-30.
In this study, the effect of farm, time, season and health was evaluated on the composition of mare's milk sold in Flanders. The content of the analyzed components (i.e. fat, fatty acids, protein, lactoferrin, lysozyme, lactose and urea) differed significantly (p < 0.0001) between farms, at a given moment in time. Within each farm, large month-to-month variations for most milk components (p <0.01 to 0.0001) were observed. The variation over time between different farms was smaller. These findings indicate that the composition of the mare's milk consumer portions varies substantially between the different farms and also over time within each farm. Season, nutrition, udder health and worm burden are believed to contribute significantly to this variation.
Hide AbstractHiller, B., Schlörmann, W., Glei, M. & Lindhauer, M. G. (2011). Food Chemistry, 125(4), 1202-1212.
Seven different types of wheat and rye bread were analysed for colorectal health related compounds, pre and post digestion, in batch fermentation model of the human intestine. Pre digestion, higher amounts of colorectal health-related dietary fibre compounds (soluble/insoluble/total dietary fibre, arabinoxylans, β-glucans) and phytochemicals (mono-/di-phenolic acids, phytic acid, hydroxymethylfurfural) were detected in wholemeal than in refined flour types of bread, as well as in rye flour types than in wheat flour types of bread. Post digestion, faecal bacterial metabolites of colorectal health promoting (acetate/propionate/butyrate, lactate, free mono-/di-phenolic acids) and impairing (amino metabolites, bile acid metabolites) activities were found in fermentation supernatants of bread samples. All types of bread positively affected faecal bacterial metabolism; among the different types of bread, the highest stimulation of organic acid production (acetate/propionate/butyrate, lactate) and the lowest detrimental bacterial enzyme activities (β-glucuronidase, urease) were detected for wheat flour bread, whereas the strongest retardation of bacterial bile acid degradation and the strongest stimulation of phenolic acid metabolite release (phenylpropionic/phenylpropenoic acid derivatives) were induced by wholemeal rye bread. This study for the first time presents a qualitative and quantitative overview over the broad spectrum of colorectal health related compounds in high- and low-fibre types of bread, pre and post in vitro digestion, and highlights the significance of bread for the preventive nutritional intervention of colorectal cancer.
Hide Abstract